BYOM function simplefun.m (the model as explicit equations)
Syntax: Xout = simplefun(t,X0,par,c,glo)
This function calculates the output of the model system. It is linked to the script files byom_bioconc_extra.m and byom_bioconc_start.m. As input, it gets:
- t is the time vector
- X0 is a vector with the initial values for states
- par is the parameter structure
- c is the external concentration (or scenario number)
- glo is the structure with information (normally global)
Note: glo is now an input to this function rather than a global. This makes the code faster (though only very little when using simplefun).
Time t is handed over as a vector, and scenario name c as single number, by call_deri.m (you do not have to use them in this function). Output Xout (as matrix) provides the output for each state at each t.
- Author: Tjalling Jager
- Date: November 2021
- Web support: http://www.debtox.info/byom.html
- Back to index walkthrough_byom.html
Copyright (c) 2012-2021, Tjalling Jager, all rights reserved. This source code is licensed under the MIT-style license found in the LICENSE.txt file in the root directory of BYOM.
Contents
Start
Note that, in this example, variable c (the scenario identifier) is not used in this function. The treatments differ in their initial exposure concentration that is set in X0mat. Also, the structure glo is not used here.
function Xout = simplefun(t,X0,par,c,glo)
Unpack initial states
The state variables enter this function in the vector _X_0.
Cw0 = X0(1); % state 1 is the external concentration at t=0 Ci0 = X0(2); % state 2 is the internal concentration at t=0
Unpack parameters
The parameters enter this function in the structure par. The names in the structure are the same as those defined in the byom script file. The 1 between parentheses is needed as each parameter has 5 associated values.
kd = par.kd(1); % degradation rate constant, d-1 ke = par.ke(1); % elimination rate constant, d-1 Piw = par.Piw(1); % bioconcentration factor, L/kg Ct = par.Ct(1); % threshold external concentration that stops degradation, mg/L
Calculate the model output
This is the actual model, specified as explicit function(s):
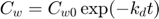
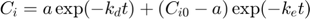
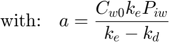
Note: this function is not completely identical to the solution of the system of ODE's. Here, I ignore the stop in degradation, which would require quite some fiddling around. Watch out that this solution is only valid when ke is NOT equal to kd!
Cw = Cw0 * exp(-kd*t); % concentration in water a = Cw0 * ke * Piw/(ke-kd); % this factor occurs twice in the solution Ci = a*exp(-kd*t) + (Ci0 - a) * exp(-ke*t); % internal concentration Xout = [Cw Ci]; % combine them into a matrix